(In collaboration with Dmitri Vezenov’s group, Chemistry Department, Lehigh University)
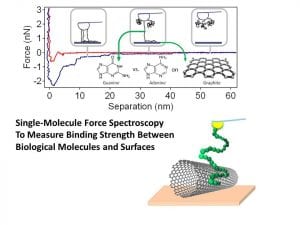
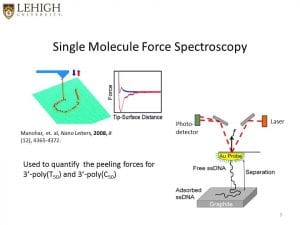
We use single-molecule force spectroscopy, that is, using an atomic force microscope tip to adsorb and then to forcibly desorb molecules from surfaces, for example, DNA off graphite or carbon nanotubes.
See also recent papers with S. Iliafar as first author. Here are some pictures that explain the process a bit.
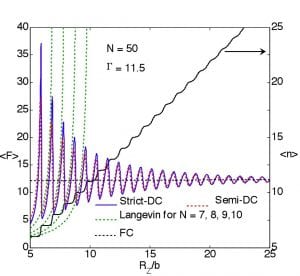
We have analyzed the statistical thermodynamics of peeling single-stranded DNA (ssDNA) from the surface of graphite. Using recently measured parameters, we model ssDNA as a freely jointed chain strongly adsorbed to a frictionless substrate. Under force control, we have obtained several exact closed-form results that, in agreement with single-molecule experiments, predict peeling under a steady force and provide a relation between this force and the underlying adhesion free energy. During peeling under displacement control we predict that, for finite length chains (< 25 bases), the equilibrium response will display spikes in force that decrease in magnitude with increasing end-to-end distance of the desorbed chain. These force spikes carry information about the underlying sequence of ssDNA, which might thus be measurable with a sufficiently stiff loading system.
(Manohar and Jagota, PRE 2010)