Marcos Pires is an Assistant Professor of Chemistry at Lehigh. Below he discusses some of current work of his lab.
In many instances, human pathology can be directly tied to the proteins that reside out the outside, on the surface, and within human cells. This in itself should not be surprising. After all, proteins are well established as acting as the workhorses of cells. Not only are proteins responsible for a high volume of chemical transformation that need to take place for cells to remain viable, they also have a series of additional roles in structuring biomacromolecules, inter- and intra-cellular communication, and storage. Through all these functions, diversity within the protein matrix becomes essential. After all, there are only 20-22k genes that encode human proteins in the human genome. The question that naturally arises is: how do human cells perform all the necessary and incredibly diverse tasks with such a limited set of building block?
During the past decade, intensive research in this area has demonstrated that following their biosynthesis, the structure of proteins is only an initial template. Proteins can be thought of as strings (of variable lengths) made with 20 building blocks. These building blocks are amino acids. All proteins in nature are made from the same 20 building blocks. Depending on one’s view, 20 building blocks to make all the proteins in the natural world (from complex human cells down to microbes) may seem like a small and insufficient number. In some ways, this may be true. One possible way to expand on the make-up of these building blocks may be to change them after the string has been assembled. And this is exactly what happens to many human cells. Following their assembly out of the biosynthetic machinery (ribosomes), they are heavily decorated with a number of chemical modifications. These modifications (or post-translational modifications) serve to diversify both the structure and function of proteins. Due to their role in controlling protein function, the aberrant modification of proteins has been heavily implicated in a number of human diseases.
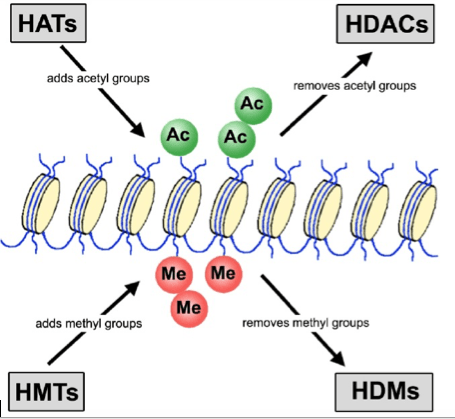
My research group is interested in discovering modifications that were not previously described in this area. We rely on fundamental chemical principles, first and foremost, to evaluate amino acid modifications that have been discovered previously. Next, we predict what combinations of chemical modifications could be tolerated based on the chemistry involved. From this analysis, we concluded that two well known modifications (methylation and acetylation) should be tolerated within the same amino acid. In other words, it may be possible that proteins are getting doubly modified at the same site via two unique pathways. By mapping this out, we predict that we may unmask a previously unappreciated signal mode within cells. Furthermore, these modifications may play roles in the development and progression of human diseases.